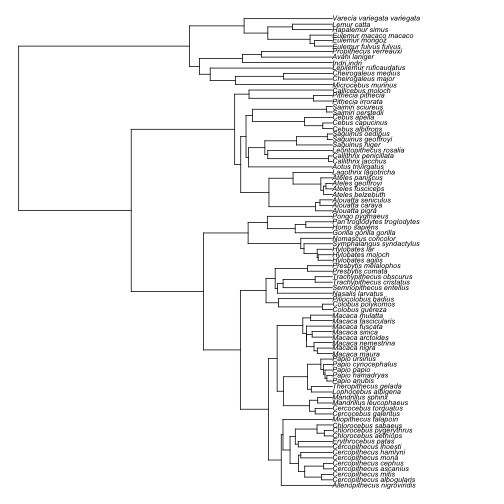
plot(tree, cex = 0.6, no.margin = T)
"ape" (Paradis et al 2004)
brain size and body size on primates ("primate_ind.txt", a tab separated text file), with more than one observation for the majority of species, individual-specific data
phylogeny ("primate_tree.phy", in phylip format) is taken from 10kTrees (Arnold et al 2010) and tailored to the data at hand
To obtain species-specific data (means and variances) from individual-specific data
data.ind = read.table("primate_ind.txt", sep = "\t", header = TRUE) body_mean = tapply(data.ind$body, data.ind$species, mean) brain_mean = tapply(data.ind$brain, data.ind$species, mean) body_N = tapply(data.ind$body, data.ind$species, length) brain_N = tapply(data.ind$brain, data.ind$species, length) body_var = tapply(data.ind$body, data.ind$species, var) brain_var = tapply(data.ind$brain, data.ind$species, var) se <- function(x) sqrt(var(x)/length(x)) body_se <- tapply(data.ind$body, data.ind$species, se) brain_se <- tapply(data.ind$brain, data.ind$species, se)
To calculate measurement errors based on pooled CV
body_cv = sqrt(body_var)/body_mean body_cv.pool = sum(body_cv * body_N, na.rm = T)/sum(body_N, na.rm = T) body_se.adj = (body_cv.pool * body_mean)/sqrt(body_N) body_var.adj = (body_cv.pool * body_mean)^2 brain_cv = sqrt(brain_var)/brain_mean brain_cv.pool = sum(brain_cv * brain_N, na.rm = T)/sum(brain_N, na.rm = T) brain_se.adj = (brain_cv.pool * brain_mean)/sqrt(brain_N) brain_var.adj = (brain_cv.pool * brain_mean)^2
To log-transform data
lg.body_var.adj = log(body_var.adj/(body_mean^2) + 1) # note that this gives the same value for all species use the following # instead: lg.body_var.adj = log(body_se.adj^2/(body_mean^2) + 1) lg.body = log(body_mean) - 0.5 * lg.body_var.adj lg.brain_var.adj = log(brain_se.adj^2/(brain_mean^2) + 1) lg.brain = log(brain_mean) - 0.5 * lg.brain_var.adj
To create data frame at the species level
data.spec = data.frame(row.names = 1:length(body_mean), species = row.names(body_mean), cbind(body_mean, body_N, body_var, lg.body, lg.body_var.adj, brain_mean, brain_N, brain_var, lg.brain, lg.brain_var.adj), check.rows = TRUE, check.names = TRUE)
To import and check phylogeny:
library(ape) tree = read.tree("primate_tree.phy") str(tree)
## List of 4 ## $ edge : int [1:170, 1:2] 87 88 89 90 91 92 92 93 94 95 ... ## $ Nnode : int 85 ## $ tip.label : chr [1:86] "Allenopithecus_nigroviridis" "Cercopithecus_albogularis" "Cercopithecus_mitis" "Cercopithecus_ascanius" ... ## $ edge.length: num [1:170] 26.19 16.81 8.59 6.54 2.98 ... ## - attr(*, "class")= chr "phylo" ## - attr(*, "order")= chr "cladewise"
To plot phylogeny:
plot(tree, cex = 0.6, no.margin = T)
